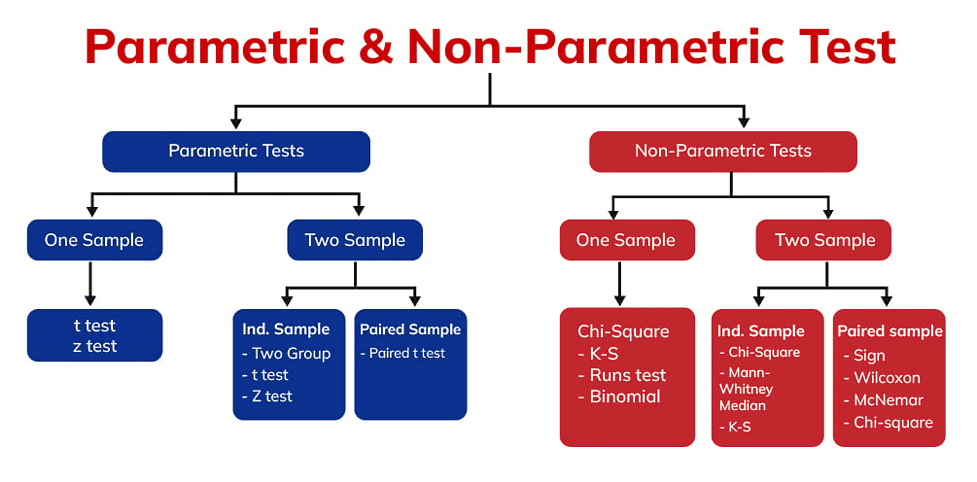
Stem cell research examines everything from gene expression to differentiation capacities to therapeutic potentials. With such diverse data types and complex experiments, researchers must determine the most appropriate statistical tests to analyze results. Parametric and nonparametric methods both play essential roles in stem cell data analysis.
Parametric tests make assumptions about population parameters and rely on normally distributed data. Tests like ANOVA, t-tests, and Pearson correlation are standard parametric methods used to compare means, determine differences between groups, and assess linear relationships. In stem cell studies, they can evaluate outcomes like gene expression, cell counts, quantitative PCR data, and other continuous biological measurements.
Parametric tests and analogous nonparametric procedures
However, the assumptions of parametric tests often need to be met in stem cell research. Data frequently violates normality due to small sample sizes, skewed distributions from zero-bounded data like cell counts, and the complexity of biological systems. Transformations may improve normality in some cases. If not, nonparametric methods are more appropriate and robust.
Nonparametric tests do not assume normality or make distributional assumptions. They compare medians rather than means and look at rank order rather than exact values. Examples include the Mann-Whitney U, Wilcoxon signed rank, Spearman correlation, and Kruskal-Wallis tests. These are useful in stem cell work for comparing two groups, paired data, correlation, and multiple group comparisons with ordinal or continuous data.
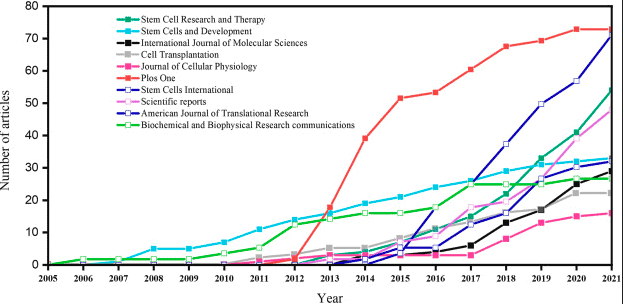
Researchers may measure stem cell data on ordinal scales, such as stages of differentiation or subjective grading scales. Standard nonparametric tests cannot be applied directly to ordinal data, but extensions like the Jonckheere-Terpstra and Cochran-Armitage tests allow for analyzing trends in ordinal data.
Another consideration is paired versus unpaired data. Parametric tests like t-tests have paired and unpaired versions. Nonparametric tests also have this distinction – Wilcoxon signed rank for paired and Mann-Whitney U for independent data. Research design determines which version to apply in stem cell therapy for metabolic diseases.
Modern stem cell experiments often involve high throughput techniques like RNA sequencing, which generates count data. This count nature lends to Poisson or negative binomial distributions rather than normality. Generalized linear models are an alternative for modeling count outcomes in cell biology.
Multi-factor studies require special attention to interactions and nested designs. Two-way ANOVA is used for factorial parametric analyses, but the rank-based Scheirer-Ray-Hare test can handle non-normal data with two factors. For nested data, mixed models or Kruskal-Wallis are suitable nonparametric approaches.
Parametric and Nonparametric: Demystifying the Terms
In all cases, graphical examination and statistical testing of distributional assumptions should precede analysis to determine parametric or nonparametric appropriateness. Stem cell scientists must remain flexible in selecting the most valid statistical tests for each experiment based on data characteristics.
While parametric methods have more power when their assumptions are met, nonparametric procedures will provide legitimate results for non-normal data. Parametric models can be more sensitive for detecting small effect sizes with adequate samples. However, with limited stem cell samples, which is standard, nonparametric methods often yield more robust conclusions.
Ideally, parametric and nonparametric techniques are applied to evaluate stem cell data from multiple perspectives thoroughly. This approach takes advantage of the breadth of modern statistical tools to produce meaningful results that withstand scrutiny since stem cell therapies often eventually progress to human clinical trials. Rigorous design and analysis are crucial to maximizing stem cell research potential.